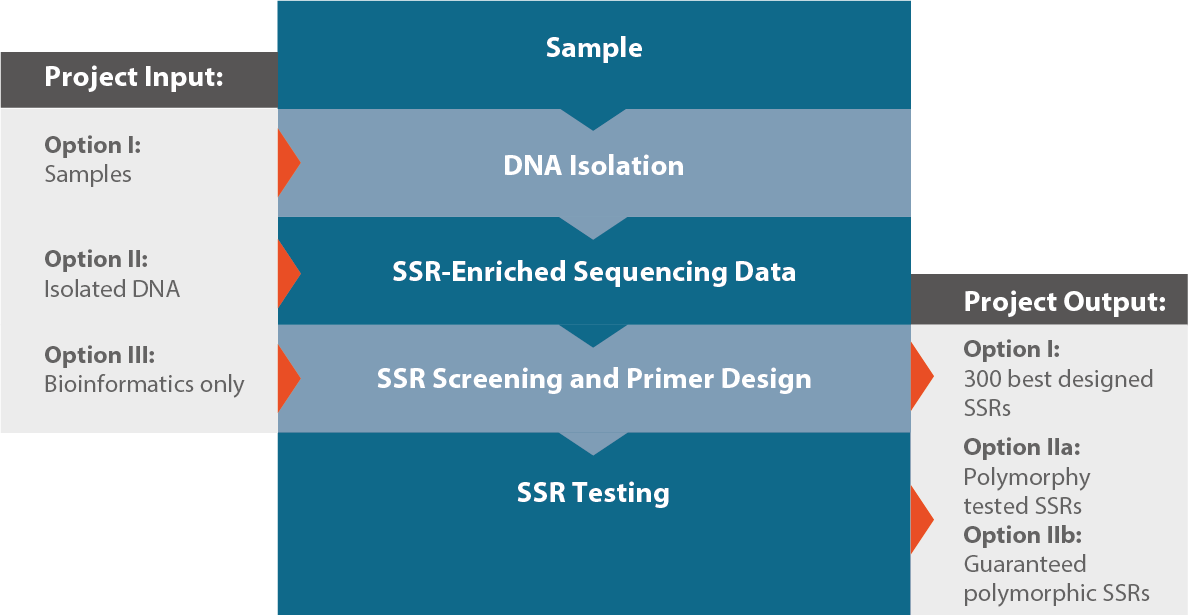
Genome-wide simple sequence repeats (SSR) markers discovered from whole-genome sequence comparisons of multiple spinach accessions | Scientific Reports
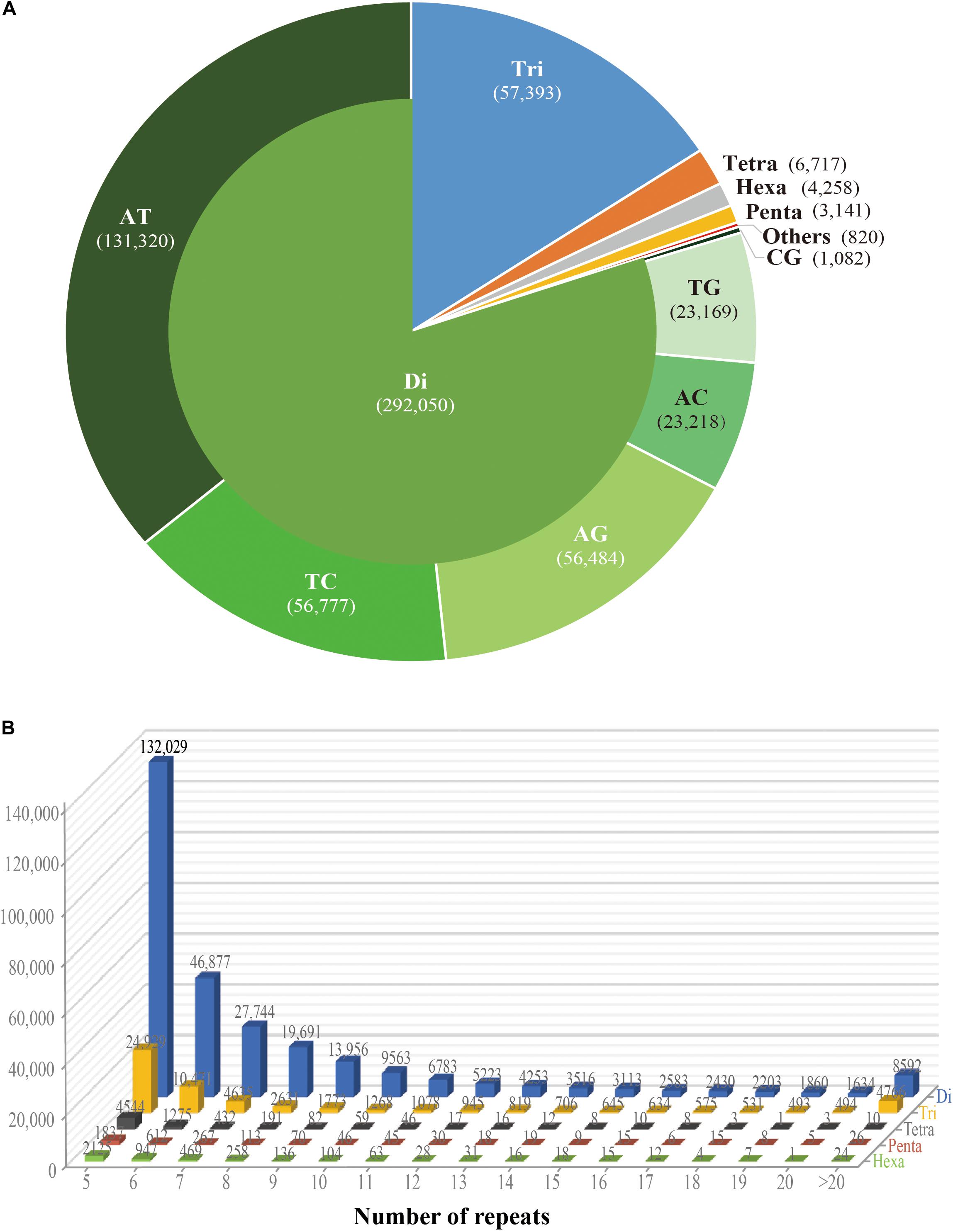
Frontiers | Development of a Large Gene-Associated SSR Marker Set and in-Depth Genetic Characterization in Scarlet Sage
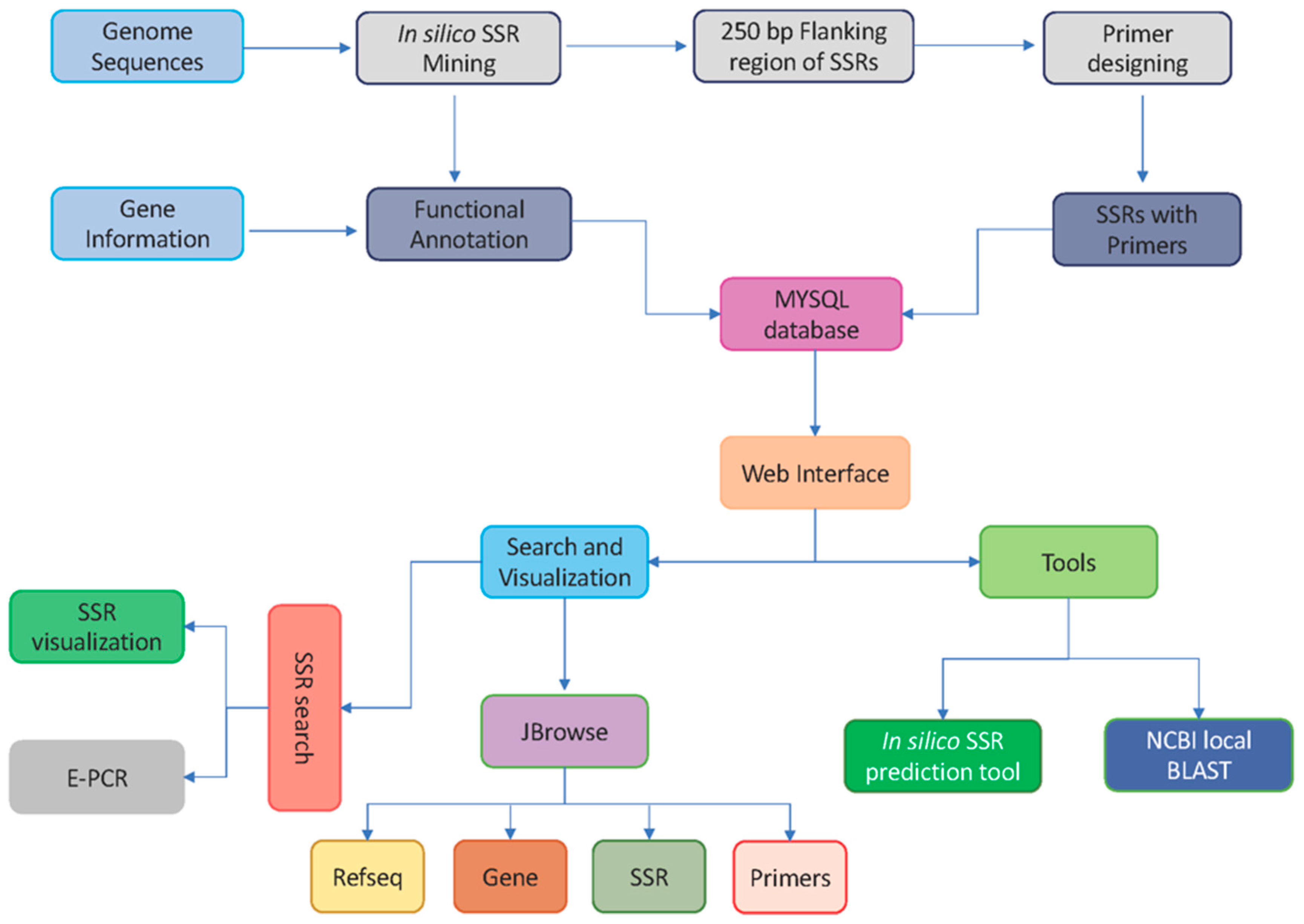
Genes | Free Full-Text | citSATdb: Genome-Wide Simple Sequence Repeat (SSR) Marker Database of Citrus Species for Germplasm Characterization and Crop Improvement
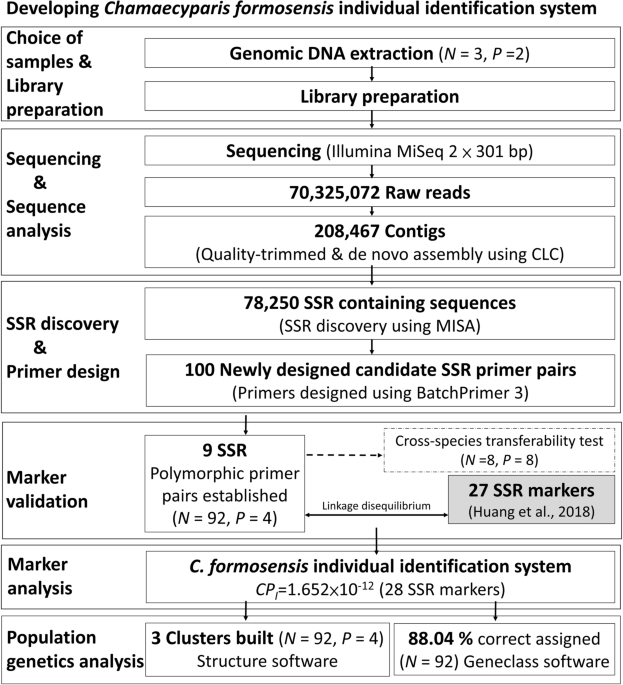
SSR individual identification system construction and population genetics analysis for Chamaecyparis formosensis | Scientific Reports

Molecules | Free Full-Text | Mining and Development of Novel SSR Markers Using Next Generation Sequencing (NGS) Data in Plants

Cross-species transferability of genomic SSR markers and genetic diversity among Asparagus racemosus Willd. Accessions - ScienceDirect

Forests | Free Full-Text | Genetic Diversity and Association Analysis among Germplasms of Diospyros kaki in Zhejiang Province Based on SSR Markers
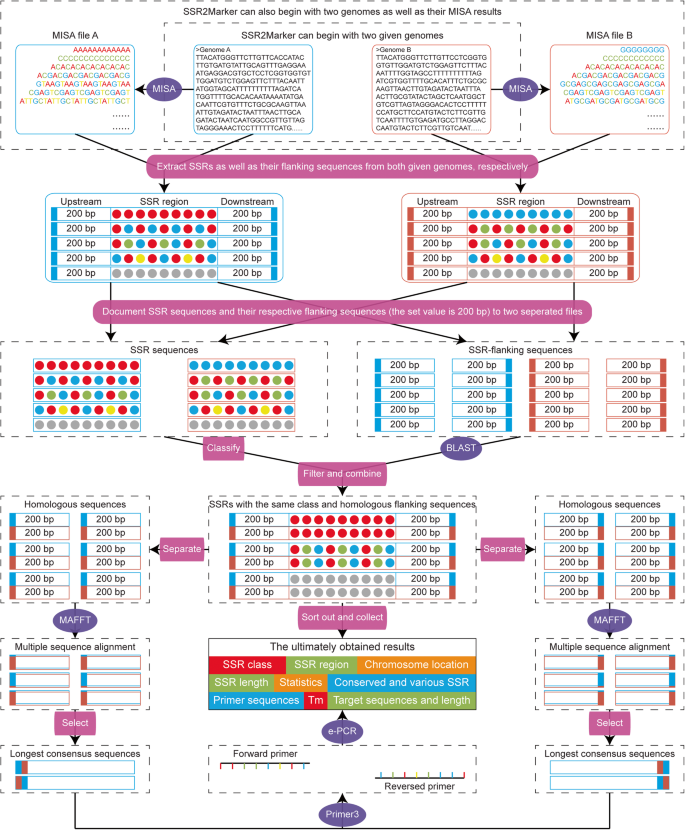
SSR2Marker: an integrated pipeline for identification of SSR markers within any two given genome-scale sequences | Molecular Horticulture | Full Text
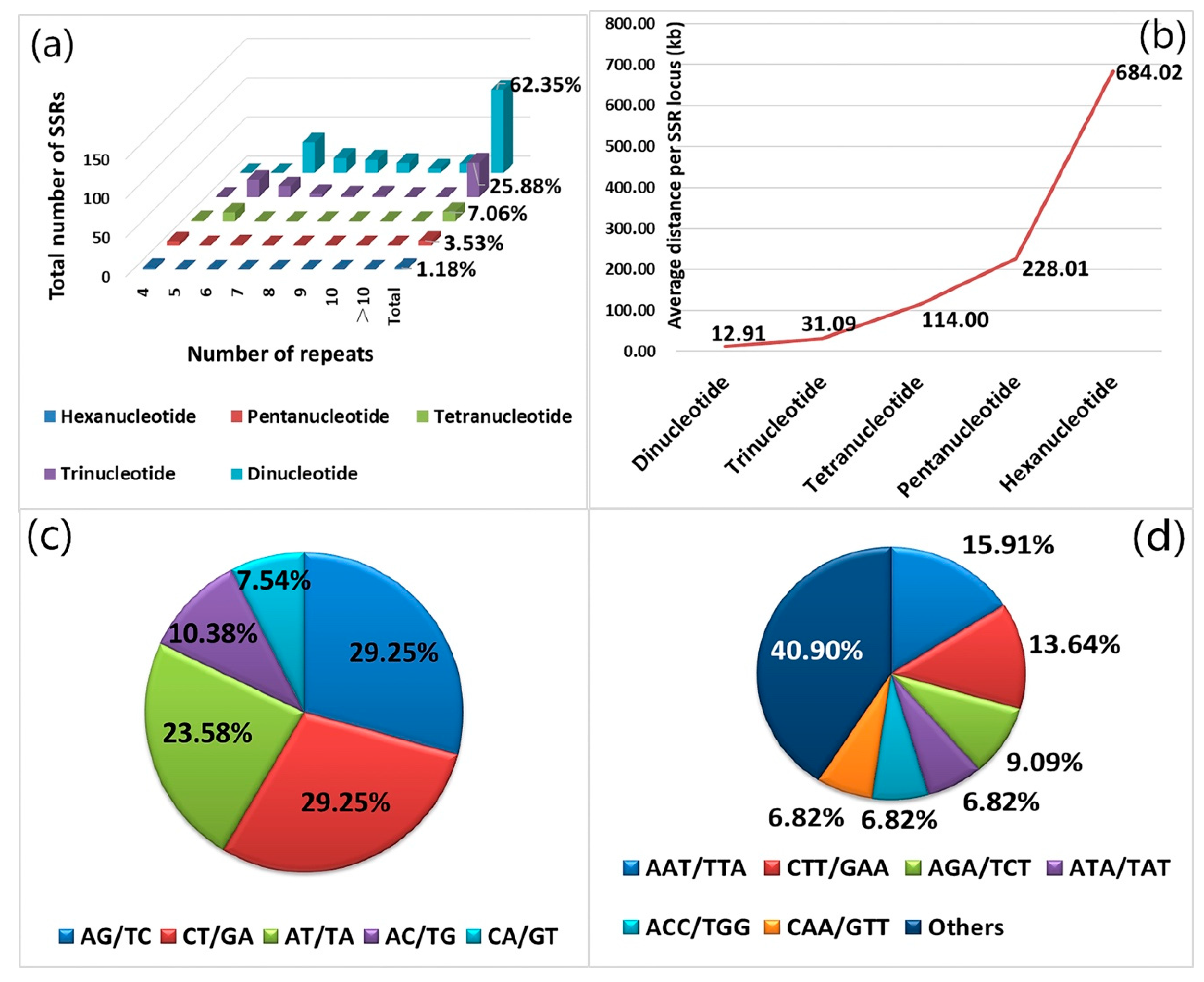
Forests | Free Full-Text | Development and Application of EST-SSR Markers for DNA Fingerprinting and Genetic Diversity Analysis of the Main Cultivars of Black Locust (Robinia pseudoacacia L.) in China
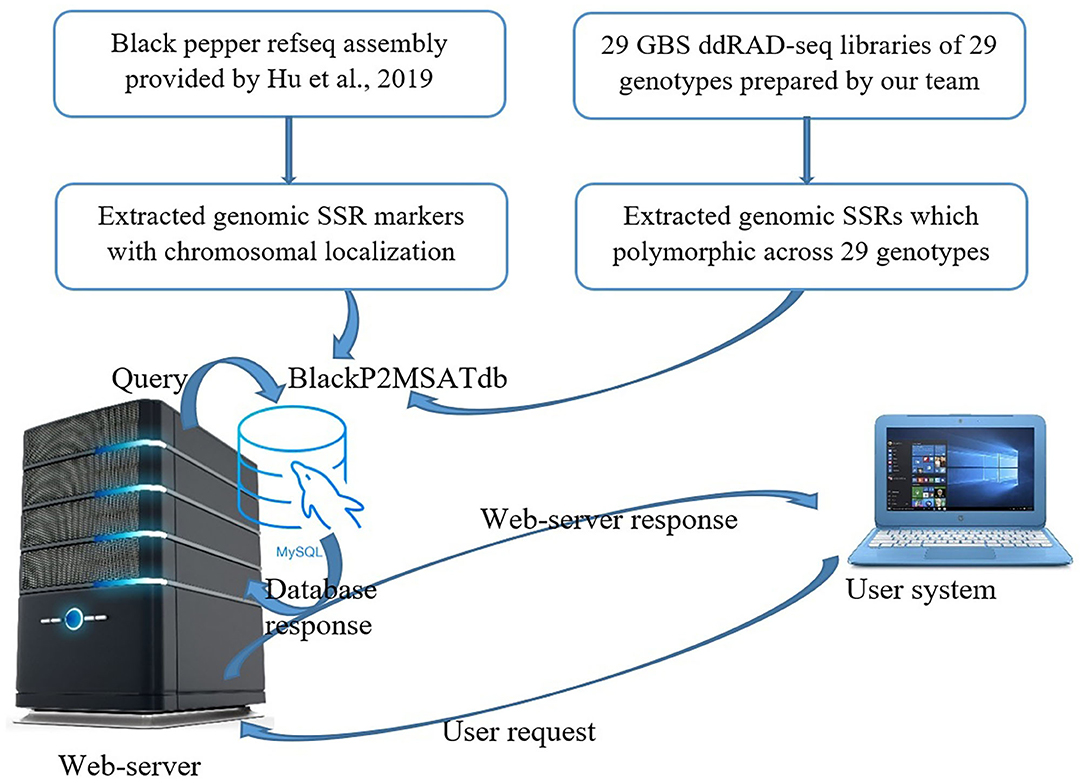
Frontiers | Rapid Genome-Wide Location-Specific Polymorphic SSR Marker Discovery in Black Pepper by GBS Approach
Genome-Wide Characterization of Simple Sequence Repeat (SSR) Loci in Chinese Jujube and Jujube SSR Primer Transferability | PLOS ONE